The BIGSdb website Policy concerning the platform & data use agreement and the privacy notice of BIGSdb-Pasteur was updated on March 25, 2024. Please consult it before using the platform and the data. If any questions, contact us.
Getting an account
If you do not have an account yet, please create a new account through the web site Register for a site-wide account and next register your account with the database(s) in which you wish to submit data via the automated submission system. If you already have an account associated with a database, you can reset your password, update your profile and register your account with specific databases in the site-wide account settings page.
Please ensure that you enter a valid E-mail address, as account validation details will be sent to this address. Please fill the box with your complete details using proper first letter capitalization of names and full affiliation details, i.e. department, institution, country (avoiding acronyms). This information will appear with any data that you submit.
Submissions
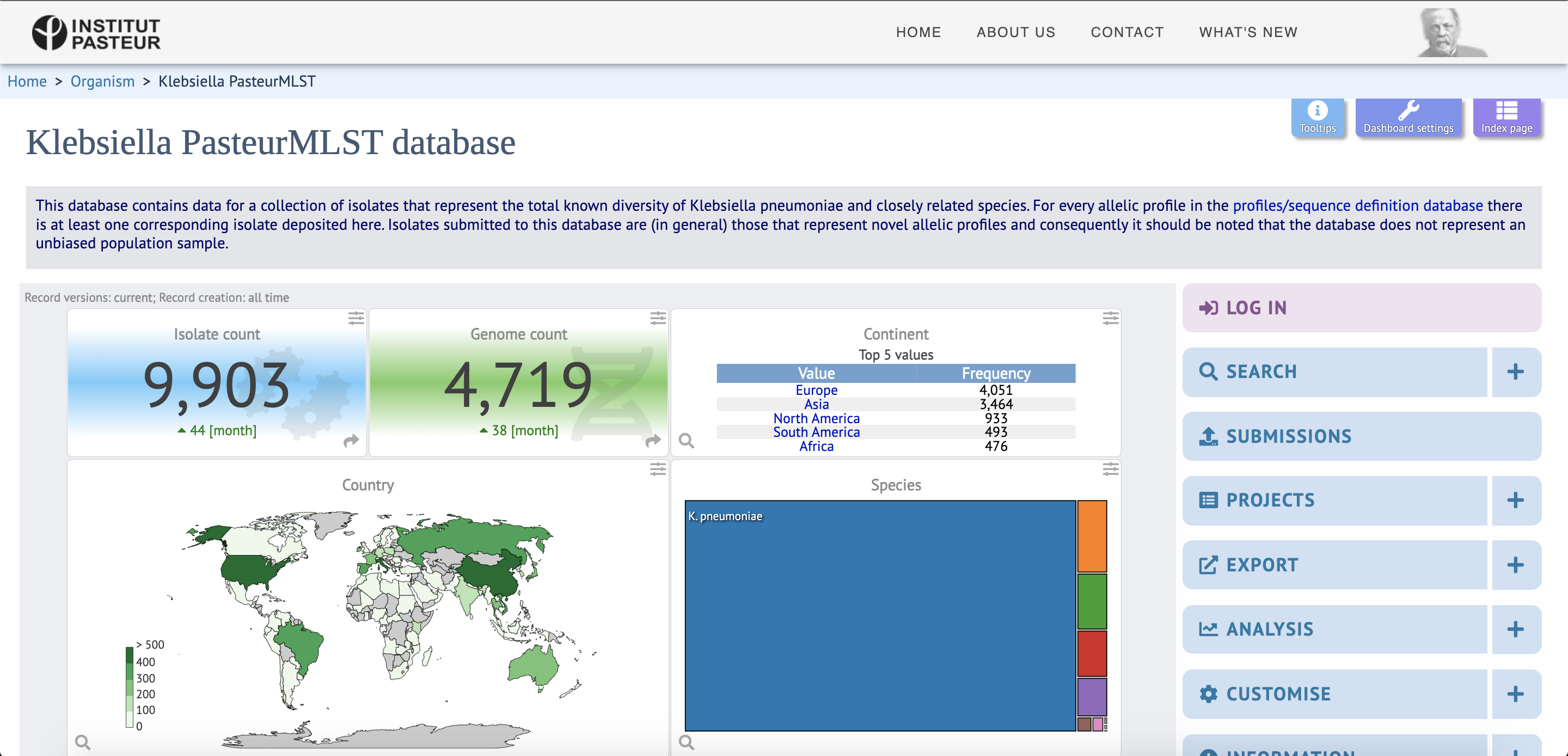
Please submit all data (isolate’s records and sequence data) for allele and profiles definitions through your BIGSdb-Pasteur submitter account. Once your submission is completed, you and the curators will automatically receive a notification. For more information about how BIGSdb-Pasteur handles your data please read our Policy.
Genomes (RECOMMENDED):
Users are requested to submit only high-quality sequences, generated from pure cultures and sequenced at a minimum coverage of 40X, that comply with the KlebNET-GSP quality criteria. Assembly files consisting of highly fragmented contigs (> 500 contigs or N50 < 20K) or presenting a cumulative contigs length outside the typical range of the Kp species (4.5-6.5Mb) will not be accepted. Submissions containing low quality assemblies may be entirely rejected.
NB. We no longer support designation of new alleles and new STs based on Ion Torrent (nor Roche/454) sequencing, although assemblies generated from such technolgies can be tagged for known alleles. We recommend you submit only assemblies either generated from Illumina short-reads or combining both short and long reads (hybrid assemblies).
Once logged into the Isolates and genomes database, click on SUBMISSIONS to access the submission panel. Choose genomes to provide the isolate information (use the template available there) and assembly files (closed or draft genomes in FASTA format). A guidance note is available in the headers of the template to help you to fill-out the metadata fields. Please label your assembly files exactly as the assembly_filename of the template and in accordance to the isolate name. For example, if the isolate name is ATCC13883 in the template, the assembly file should be ATCC13883.fas. The suffixes .fa, .fasta or .fna are also acceptable.
NB. When genomes are provided, users are not requested to also submit the novel allele(s) and profile(s) in the definition database; curators will perform MLST alleles and profiles curation by default. In addition, users can ask to design further schemes (see examples of wzi, cgMLST, AbST, YbST, ...) using the Messages box of the submission page.**
The submitted genomes can be embargoed. If you wish your genomic sequences to become public at a later stage, for example, when a publication is accepted (or no later than one year after submission, typically); please indicate this via the Messages box of the submission page.
MLST profiles:
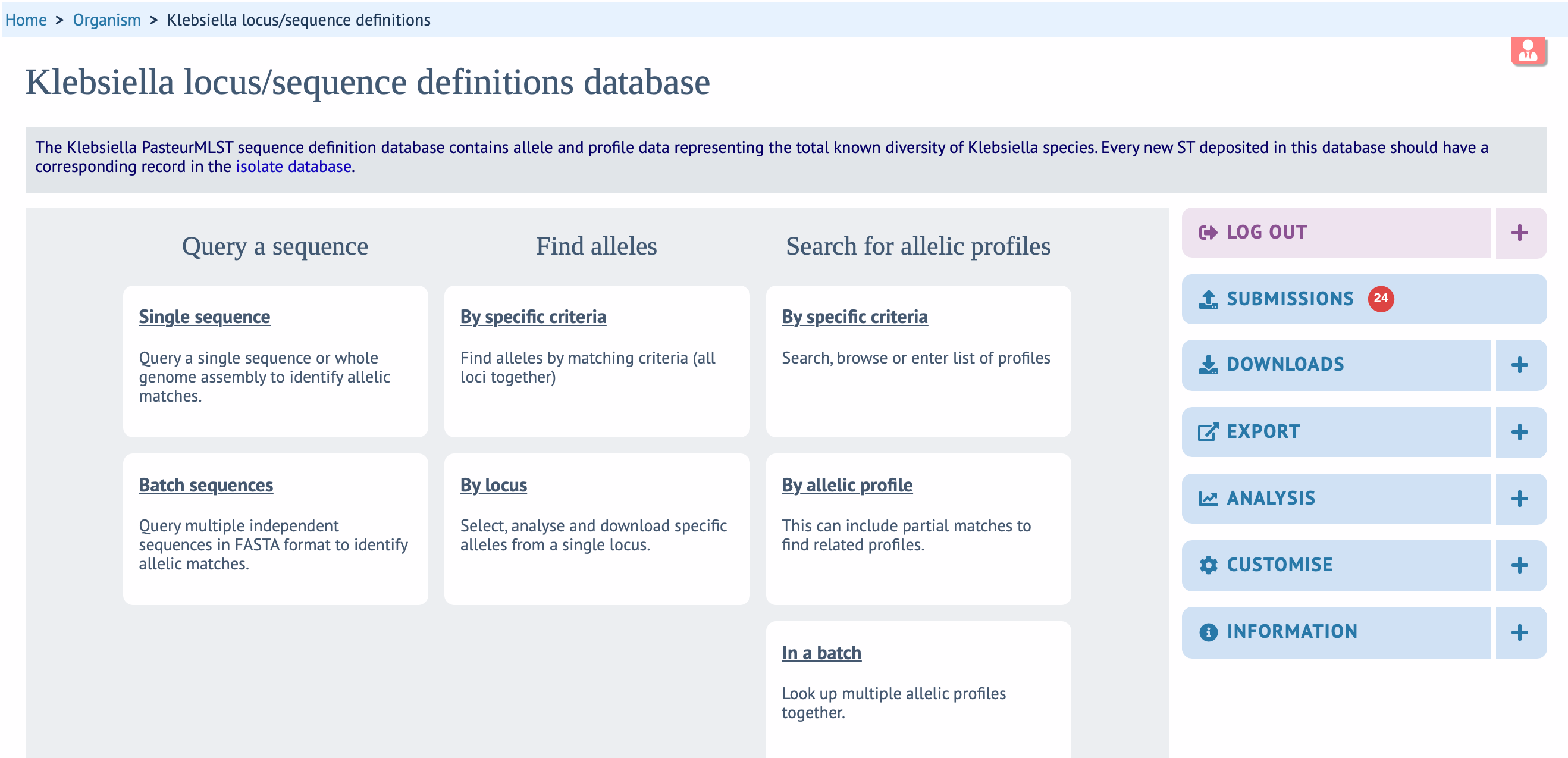
For each novel profile, users are requested to submit at least one isolate record (with minimal metadata required) and, if available, related genome assembly through the Isolates and genomes database. Multiple novel profiles can be provided as a single submission.
Please see the Genomes (RECOMMENDED) procedure above or follow the procedure below if genomes are not available:
- Log into the Isolates database, click on SUBMISSIONS and choose isolates to provide the isolate records (use the template available there). You can submit multiple isolates in a single submission.
- Log into the Alleles and profiles database, click on SUBMISSIONS and choose MLST profiles to provide the profiles (use the template available there). You can submit multiple MLST profiles in a single submission. Please label the novel profile as the related isolate submitted in the Isolates database.
NB. The submitted profile should correspond to results obtained querying your sequence into BIGSdb-Pasteur Alleles and profiles database, as this database may be more up to date than external databases that download and provide our curated data on other platforms.**
Alleles:
For allele(s) extracted by genome assemblies, users are requested to follow the Genomes (RECOMMENDED) procedure above.
For allele(s) assembled from trace files, please follow the procedure below:
Once logged into the Alleles and profiles database, click on SUBMISSIONS, choose alleles to submit any number of new sequences for a single locus and related trace files. Sequences should be trimmed to the correct start/end sites for the selected locus. Trace files should be labeled starting with the strain code (avoiding special characters such as space, dot, slash, etc.), followed by underscore, followed by the gene name written exactly as in the database (e.g. gapA), followed by underscore, followed by any information you would like, followed by .ab1 or .scf. Example: SB1_gapA_398509385_F.ab1 and SB1_gapA_398509385_R.ab1.
Data release
We typically process submissions within 48h. Once the curation is complete, allele definitions and profiles will be released immediately, as they form the basis for the public nomenclature of genotypes. You will be notified and asked to control if all data integrated in the BIGSdb database are correct.
We encourage the submission of all isolates of your studies, so that the database would be as representative of the natural populations as possible. Thus, please do not restrict yourself to submitting only isolates that represent new profiles: isolates with already known MLST types are valuable as well.
We also encourage immediate release of your isolates provenance data and genomic sequences. However, in the case you would like to ask for an embargo period (e.g. prior to publication), please specify it to the curators in the Messages box of the submission page or mailing curators. Data release should ideally be when your publication is accepted or no later than one year after submission to our database. Curators may automatically submit your data to the public database after this period.
We kindly ask you to inform us in the case of data publication or metadata rectifications to allow us to keep current records updated.
Acknowledgments
Curation and maintenance of BIGSdb-Pasteur databases is performed on a voluntary basis by a handful of colleagues, who dedicate their time to the community.
We appreciate if you can recognize our efforts in the acknowledgments section of your publications:
We thank the Institut Pasteur teams for the curation and maintenance of BIGSdb-Pasteur databases at http://bigsdb.pasteur.fr/.
References
Please quote the original publications on the development of nomenclature schemes, which are indicated on the references page.
Collaboration
For submissions of large amounts of data, or when you would like to request further analyses in addition to curation, we may ask you to add the person of the curator team in charge of analyzing your data among your co-authors.